MSPAC
Contents
Modified Spatial Autocorrelation (MSPAC) Toolbox - Overview
After downloading the tutorial database and viewing the group of component waveforms in the data viewer, open the MSPAC toolbox by pushing the following plugin icon ![]() . Alternatively you can drag your group directly into the MSPAC toolbox icon. The MSPAC toolbox is now attached to the signal viewer.
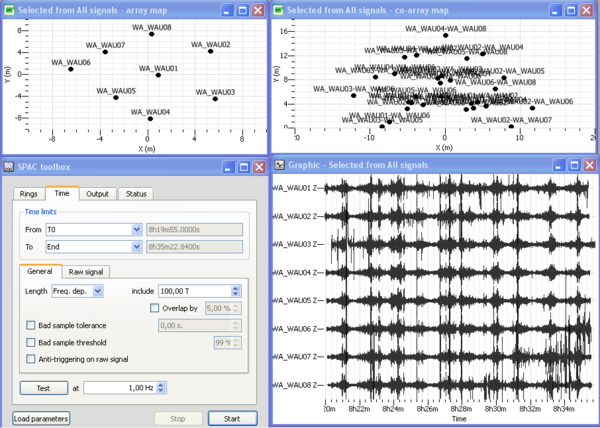
Four windows should then open as displayed in the figure on the right: (1) a signal time viewer, (2) a map displaying array station locations, (3) a map displaying co-array stations location and (4) the SPAC processing toolbox. This processing toolbox is composed of four tabs:
. Alternatively you can drag your group directly into the MSPAC toolbox icon. The MSPAC toolbox is now attached to the signal viewer.
Four windows should then open as displayed in the figure on the right: (1) a signal time viewer, (2) a map displaying array station locations, (3) a map displaying co-array stations location and (4) the SPAC processing toolbox. This processing toolbox is composed of four tabs:
- Rings tab allows to design rings from the co-array sensors map;
- Time tab allows to select the time limits and part of the signals to be processed and the time window length;
- Output tab is used to set up the frequency band to be processed together with the sampling scale type (linear, log) and number of frequencies, and the output filename
- Status tab provides information about computing status.
Note that you may process only the vertical component or the three components (see Bettig et al. (2001) [1] and Köhler et al. (2007) [2]). In both cases, spatial autocorrelation coefficients will be computed and results statistics can be displayed and edited by using max2curve command tool for the three- and the vertical components and by using spac2disp for the vertical components. However, computation of Rayleigh and Love waves phase velocities from three-component processing is not available yet. The following tutorial steps are thus only focused on vertical component processing.
Defining rings
General background regarding definition and usage of co-arrays and rings can be found here.
In Rings tab:
- Push on
 button to add a ring
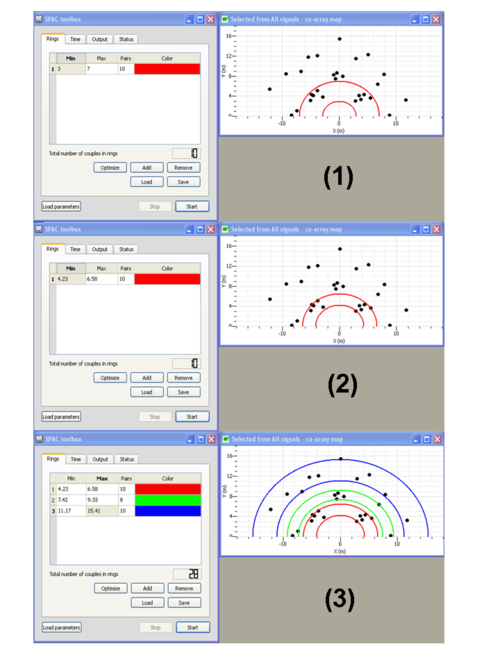
button to add a ring - Specify the inner and outer circles radii of the ring (see step (1) in the figure on the left)
- Press on
 buttom to adjust the inner and outer circles radii to fitting at best sensors pairs location (see step (2) in the figure on the left). This ring is composed of 10 pairs of sensors as specified in the Pairs column.
buttom to adjust the inner and outer circles radii to fitting at best sensors pairs location (see step (2) in the figure on the left). This ring is composed of 10 pairs of sensors as specified in the Pairs column. - Define the next rings (see step (3) in the figure on the left). Note that you can associate a specific color to each ring.
- Note that you add or remove specific ring by pushing on the
 or
or  buttons
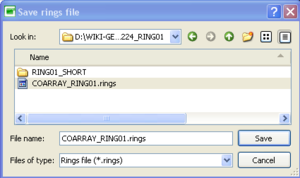
buttons - Once done, click on
 to save your rings design.
to save your rings design.
- The rings design are saved in ascii format with the following format: inner circle radius (in meters), outer circle radius (in meters), RGB color code associated to the ring. Note that any rings design can also be loaded by pushing on the
 button.
button.
# MinRadius MaxRadius Red Green Blue 4.23 6.58 255 0 0 10 7.42 9.33 0 255 0 8 11.17 15.41 0 0 255 10
Parameters settings
Time limits
- Select the following time limits to be processed.
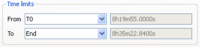
- Select a Freq. Dep. window length of 100 T. Note that the most common values are ranging from 50 to 200 T.

- By pushing on
 , you can also load the .log file containing all the input parameters saved from a previous computation.
, you can also load the .log file containing all the input parameters saved from a previous computation.
Defining the outputs and starting the computation
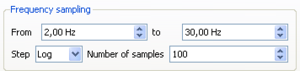
- Choose 2.0 and 30 as the minimum and the maximum center frequencies for processing.
- Specify the number of frequencies (100) and the sampling scale type (constant on a logarithmic scale).
- More details about the meaning of these options can be found in Sampling Frequency page.
There are two ways for saving computation results:
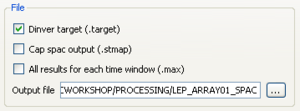
- check the Dinver target box and provide the output file name in the editing box. The .target output file will contain only the statistics: mean, standard deviation and number of processed windows for each autocorrelation sample. This file contains also the list of selected rings. This file can be further edited by using the spac2disp tool.
- check the All results for each time window box and provide the output file name in the editing box. The .max output file will contain the autocorrelation results for each processed time window and each ring. This file can be further edited by using max2curve tool.
- Then, click on
 to start the computation. Status of the computation can be checked in the "Status" tab as displayed below.
to start the computation. Status of the computation can be checked in the "Status" tab as displayed below.
Graphical display of spatial autocorrelation curves / contents of output files
The spatial autocorrelation coefficients and the parameters used for the computation are saved in a .target (or .max) file and a .log ascii file, respectively. The .log file is saved in the same folder as the .target (.max) file and contains all parameters used for the computation. This .log file can be re-used in further computation by pushing on ![]() button in the MSPAC toolbox.
button in the MSPAC toolbox.
Graphical display of spatial autocorrelation estimates and corresponding phase velocities
The .target file is Xml file. Graphical display of this file can be done by importing the .target file in the target menu of Dinver (Importing the autocorrelation curve to fit) or by using spac2disp that can be launched either from the windows Geopsy menu either from command line.
$ spac2disp LEP_ARRAY01_SPAC.target
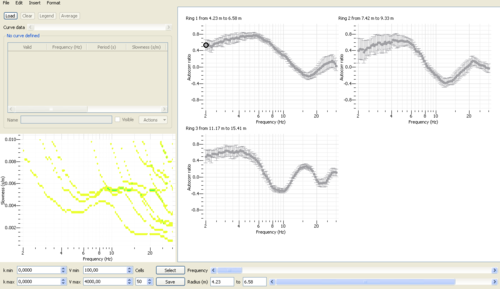
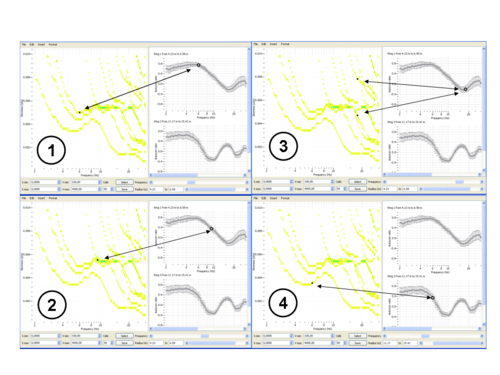
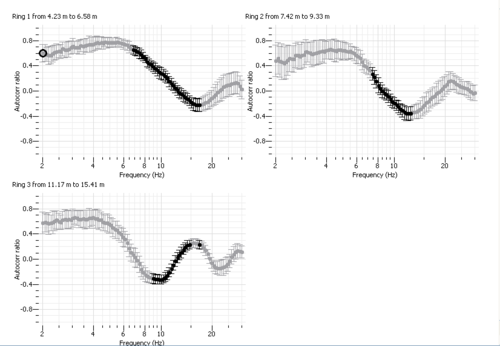
- Spatial autocorrelation curves are displayed for each ring in the right panel.
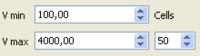
- Histograms of phase velocities derived from overallset of spatial autocorrelation values are displayed in the lower left panel. Histograms are computed within the velocity range [Vmin - Vmax] - equidistandly sampled in slowness, however - with a number of cells Cells specified in the lower panel.
- By using the Frequency and Radius scroll bars and associated sliding cursors
 , one can check for each frequency and each ring relation between autocorrelation coefficient and phase slownesses as indicated by the black dots (see figure below).
, one can check for each frequency and each ring relation between autocorrelation coefficient and phase slownesses as indicated by the black dots (see figure below).
The next step is to select those spatial autocorrelation coefficients that provide phase slownesses that contribute at best to the dispersion curve.
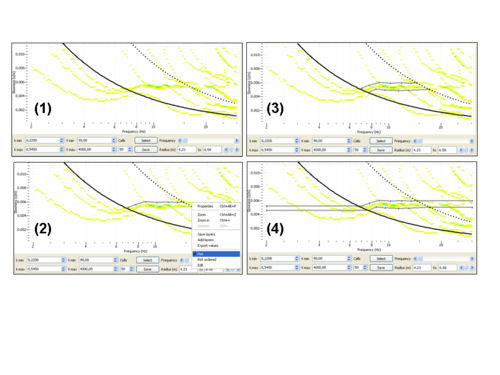
- in the lower left panel, adjust kmin and kmax
 in order to select for solutions above minimum wavenumber (kmin) and below maximum wavenumber (kmax) only (step (1) in the figure on the right). Note that these kmin and kmax do not have any special meaning and are not related to array response properties as kmin and kmax used in [[FK|FK]¨] and HRFK processing.
in order to select for solutions above minimum wavenumber (kmin) and below maximum wavenumber (kmax) only (step (1) in the figure on the right). Note that these kmin and kmax do not have any special meaning and are not related to array response properties as kmin and kmax used in [[FK|FK]¨] and HRFK processing. - right-click in the lower left panel and select the pick tab
 .
. - by clicking on the lower left panel, select the upper limit in slowness
- right-click and unselect the pick tab. See step (2) in the figure on the right.
- right-click, select again the pick tab and select the lower limit in slowness. Then unselect the pick tab. See step (3) in the figure on the right.
- once done, push the button
 in order to finalize the frequency-slowness selection (see step (4) in the figure on the right)
in order to finalize the frequency-slowness selection (see step (4) in the figure on the right) - you should then see on the right panel the spatial autocorrelation values (black dots) contributing to the selected phase slownesses as displayed below.
- the selected spatial autocorrelation estimates can now be saved in a .target file by pushing on
 button in the lower frame. This .target file can be further inverted.
button in the lower frame. This .target file can be further inverted.
- Note that the two picked curves are also displayed in the upper left frame. These curves can be edited, saved, removed, resampled, etc. You can also load a dispersion curve from other type of processing (FK, HRFK) in order to check consistency between estimates derived from different processing schemes.
Graphical display of all spatial autocorrelation estimates
The .max ASCII file contains spatial autocorrelation estimates for all processed windows, component and ring. Format of this file is the following:
- seconds from start' indicates the begin time of the processed window
- cfreq indicates (center) frequency in Hz
- icomp indicates component index (0: vertical, 1: radial, 2: transverse)
- iring indicates the ring index (from 0), in the same order as the one specified in the Rings tab of the MSPAC toolbox.
- autocorr is the spatial autocorrelation value
# File generated by Geopsy, SPAC processing # The process log is saved in file D:/WIKI-GEOPSY/DOCWORKSHOP/PROCESSING/LEP_ARRAY01_SPAC.log # icomp can take values 0 (vertical), 1 (radial), and 2 (transverse) # seconds from start | cfreq | icomp | iring | autocorr 30020 2 0 0 0.718134 30020 2 0 1 0.839546 30020 2 0 2 0.654078 30020 2.05546 0 0 0.681075 30020 2.05546 0 1 0.801923 30020 2.05546 0 2 0.710762 30070 2 0 0 0.839481
The .max file can also be edited by using max2curve tool on a command line or from the Windows Geopsy menu.
$max2curve LEP_ARRAY01_SPAC.max
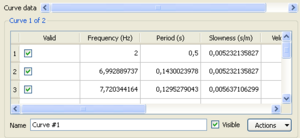
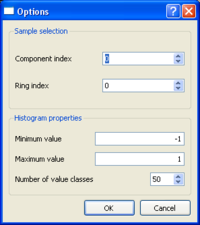
A popup window will then appear allowing to specify:
- component and ring indices as defined above
- the miminum and maximum mspac values and the number of cells to be used for defining the histogram properties
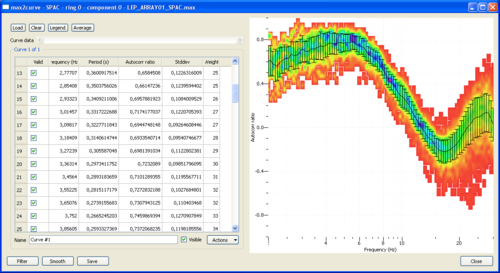
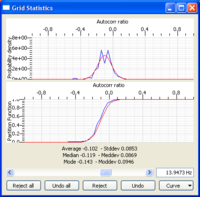
Histograms are displayed on the right panel. The color scale indicates the histogram values. By default, histograms are overlaid by the mean spatial autocorrelation +/- standard deviation curves whose corresponding values are indicated in the left panel. This 'by default' curves can be either hidden by unchecking the visible box on the bottom part of the left panel or remove by using the Remove option in the actions tab attached to the curve. Note also that you can perform several actions on this curve. Similarly to FK or HRFK, the Grid Statistics window allows you to edit the statistics in order to filter out spurious or unwanted estimates.
References
- ↑ Bettig B., P.-Y. Bard, F. Scherbaum, J. Riepl, F. Cotton, C. Cornou, D. Hatzfeld, 2001. Analysis of dense array measurements using the modified spatial auto-correlation method (SPAC). Application to Grenoble area., Boletin de Geofisica Teoria e Applicata, 42, 3-4, 281-304.
- ↑ Köhler, A., M. Ohrnberger, F. Scherbaum, M. Wathelet and C. Cornou, 2007. Assessing the reliability of the modified three-component spatial autocorrelation technique. Geophysical Journal International, Volume 168, Issue 2: 779-796. doi: 10.1111/j.1365-246X.2006.03253.x.